Project
e-OMIX offers a solution for the analysis, management and exploration of omics data through an user-friendly interface.
Metadata management and standardization.
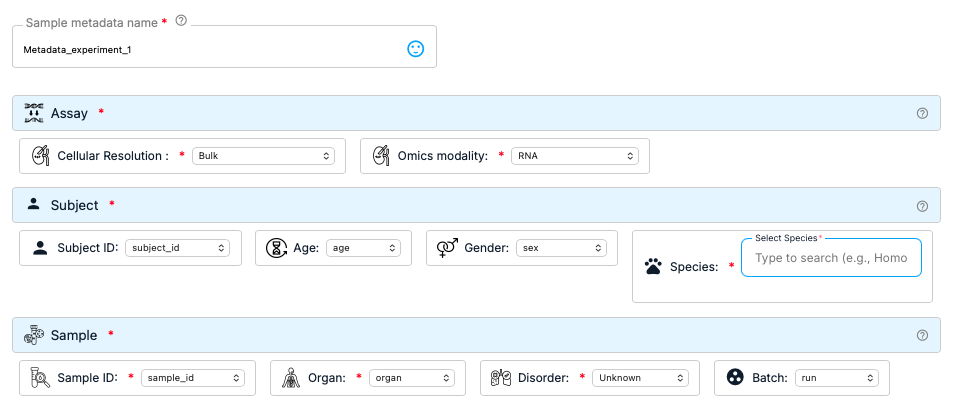
Metadata play a crucial role in omics experiments. They provide essential elements for contrasting analysis and controlling experimental variation, such as batch corrections. However, metadata lacks standardization and is often presented as flat tables with no uniform structure. e-OMIX addresses this limitation by offering a metadata standardization tool that transforms flat-table metadata into a standardized, structured format. This facilitates the exchange and comparison of experiments. Behind the scene, the metadata standardization feature leverages HL7 FHIR©, a standard format in the healthcare industry that aligns with current practices in the field.
End-to-end pipelines
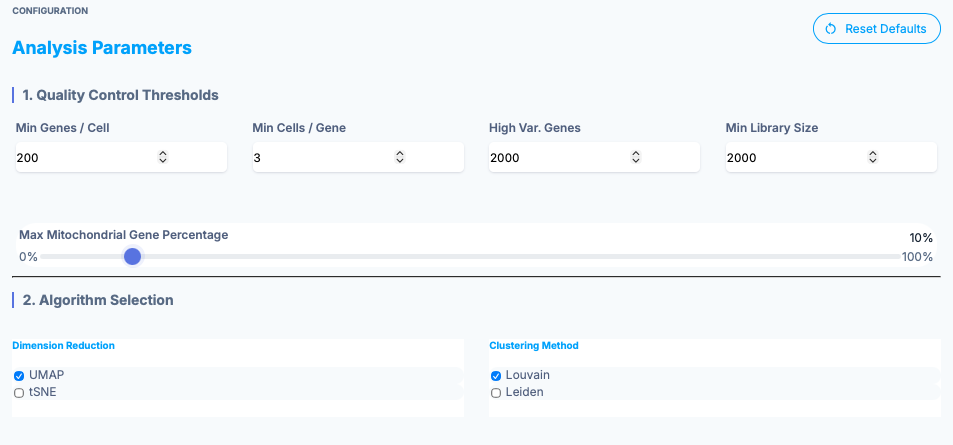
e-OMIX offers a solution for bench biologists to perform routine analyses without the need to know a programming language. Our platform provides a click-button interface to transform raw RNA sequencing results into count matrices. These matrices can then be processed with customizable filters and downloaded as R object.
Visual exploration
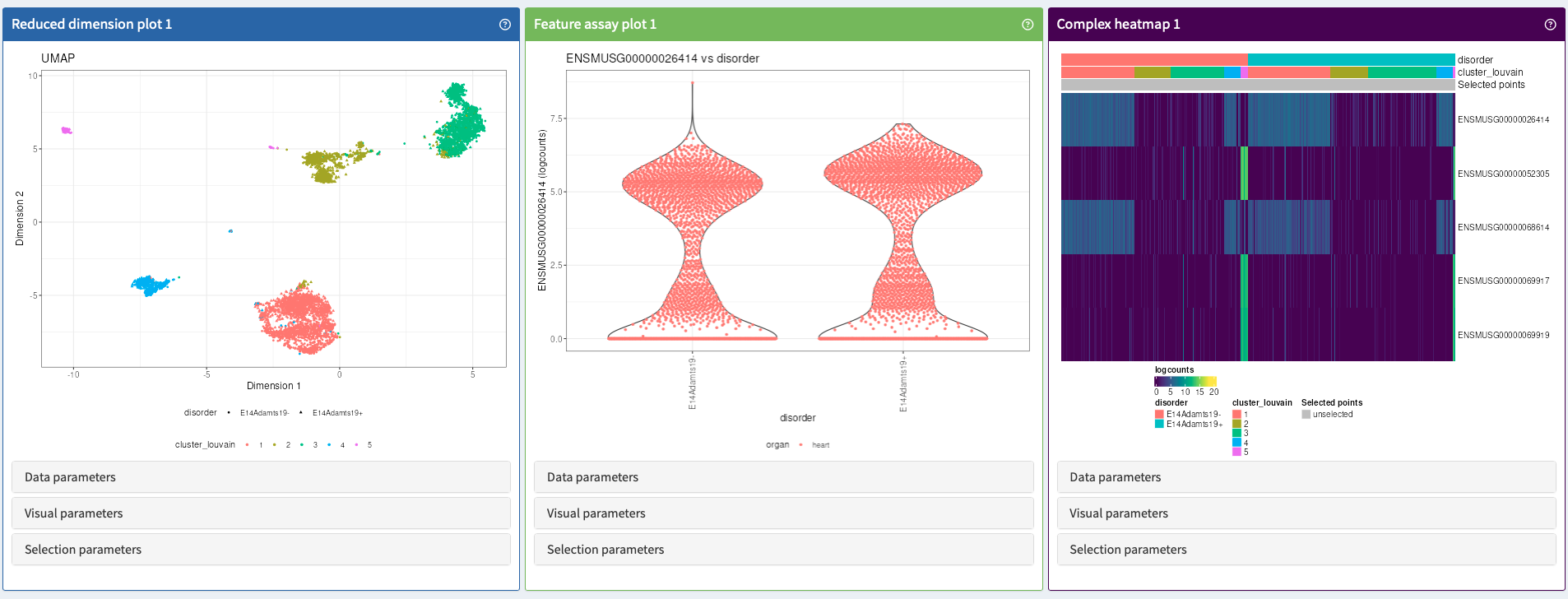
After analysis, data can be explored using a series of interactive tools that provide users with relevant biological insights. All plots can be parametrized and exported in a publication-ready format.
Download
An working version of e-OMIX can be downloaded here.
Funding
e-OMIX is part of the WAL-IMAGIN portfolio.
Financial support is provided by: